Getting Started with Powerbox¶
In [3]:
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
There are two useful classes in powerbox: the basic PowerBox,
and one for log-normal fields: LogNormalPowerBox. You can import
them like this:
In [1]:
from powerbox import PowerBox, LogNormalPowerBox
Once imported, to see all the options, just use help(PowerBox).
Create a 2D Gaussian field with power-law power-spectrum¶
For a basic 2D Gaussian field with a power-law power-spectrum, one can use the following:
In [10]:
pb = PowerBox(N=512, # Number of grid-points in the box
dim=2, # 2D box
pk = lambda k: 0.1*k**-2., # The power-spectrum
boxlength = 1.0, # Size of the box (sets the units of k in pk)
seed = 1010) # Set a seed to ensure the box looks the same every time (optional)
plt.imshow(pb.delta_x,extent=(0,1,0,1))
plt.colorbar()
plt.show()
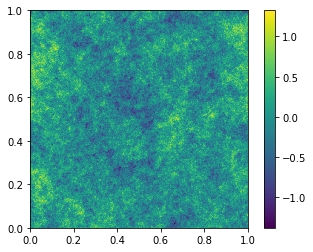
The delta_x output is always zero-mean, so it can be interpreted
as an over-density field, \(\rho(x)/\bar{\rho} -1\). The caveat to
this is that an overdensity field is physically invalid below -1. To
ensure the physical validity of the field, the option
ensure_physical can be set, which clips the field:
In [11]:
pb = PowerBox(N=512, # Number of grid-points in the box
dim=2, # 2D box
pk = lambda k: 0.1*k**-2., # The power-spectrum
boxlength = 1.0, # Size of the box (sets the units of k in pk)
seed = 1010, # Set a seed to ensure the box looks the same every time (optional)
ensure_physical=True) # ** Ensure the delta_x is a physically valid over-density **
plt.imshow(pb.delta_x,extent=(0,1,0,1))
plt.colorbar()
plt.show()
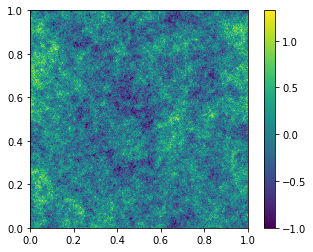
If you are actually dealing with over-densities, then this clipping solution is probably a bit hacky. What you want is a log-normal field…
Create a 2D Log-Normal field with power-law power spectrum¶
The LogNormalPowerBox class is called in exactly the same way, but
the resulting field has a log-normal pdf with the same power spectrum.
In [12]:
lnpb = LogNormalPowerBox(N=512, # Number of grid-points in the box
dim=2, # 2D box
pk = lambda k: 0.1*k**-2., # The power-spectrum
boxlength = 1.0, # Size of the box (sets the units of k in pk)
seed = 1010) # Use the same seed as our powerbox
plt.imshow(lnpb.delta_x,extent=(0,1,0,1))
plt.colorbar()
plt.show()
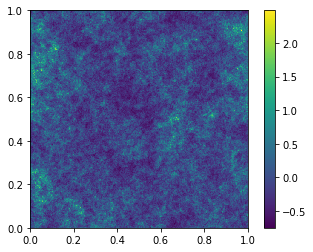
Again, the delta_x is zero-mean, but has a longer positive tail due
to the log-normal nature of the distribution. This means it is always
greater than -1, so that the over-density field is always physical.
Create some discrete samples on the field¶
powerbox lets you easily create samples that follow the field:
In [18]:
fig, ax = plt.subplots(1,2, sharex=True,sharey=True,gridspec_kw={"hspace":0}, subplot_kw={"ylim":(0,1),"xlim":(0,1)}, figsize=(10,5))
# Create a discrete sample using the PowerBox instance.
samples = pb.create_discrete_sample(nbar=50000, # nbar specifies the number density
min_at_zero=True # by default the samples are centred at 0. This shifts them to be positive.
)
ln_samples = lnpb.create_discrete_sample(nbar=50000, min_at_zero=True)
# Plot the samples
ax[0].scatter(samples[:,0],samples[:,1], alpha=0.2,s=1)
ax[1].scatter(ln_samples[:,0],ln_samples[:,1],alpha=0.2,s=1)
plt.show()
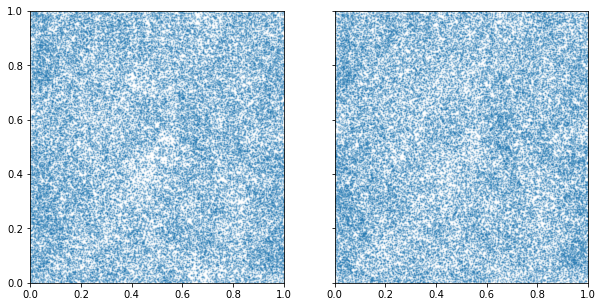

Within each grid-cell, the placement of the samples is uniformly random.
The samples can instead be placed on the cell edge by setting
randomise_in_cell to False.
Check the power-spectrum of the field¶
powerbox also contains a function for computing the (isotropic)
power-spectrum of a field. This function accepts either a box defining
the field values at every co-ordinate, or a set of discrete samples.
In the latter case, the routine returns the power spectrum of
over-densities, which matches the field that produced them. Let’s go
ahead and compute the power spectrum of our boxes, both from the samples
and from the fields themselves:
In [19]:
from powerbox import get_power
In [24]:
# Only two arguments required when passing a field
p_k_field, bins_field = get_power(pb.delta_x, pb.boxlength)
p_k_lnfield, bins_lnfield = get_power(lnpb.delta_x, lnpb.boxlength)
# The number of grid points are also required when passing the samples
p_k_samples, bins_samples = get_power(samples, pb.boxlength,N=pb.N)
p_k_lnsamples, bins_lnsamples = get_power(ln_samples, lnpb.boxlength,N=lnpb.N)
Now we can plot them all together to ensure they line up:
In [26]:
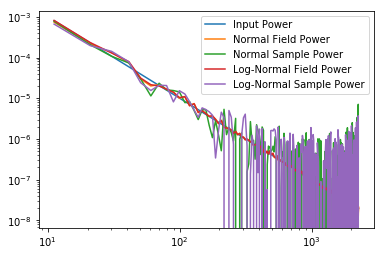
plt.plot(bins_field, 0.1*bins_field**-2., label="Input Power")
plt.plot(bins_field, p_k_field,label="Normal Field Power")
plt.plot(bins_samples, p_k_samples,label="Normal Sample Power")
plt.plot(bins_lnfield, p_k_lnfield,label="Log-Normal Field Power")
plt.plot(bins_lnsamples, p_k_lnsamples,label="Log-Normal Sample Power")
plt.legend()
plt.xscale('log')
plt.yscale('log')